- 8808 2348
- blacklord1234@gmail.com
- Mon - Sun 9:00AM - 5:00PM

Before you read on, you might want to download this entire revision notes in PDF format to print it out for your child, or yourself to read it later.
This will be delivered to your email inbox.
Compared to eukaryotes, prokaryotes are much simpler creatures. They do not have the many membrane-bound organelles that define eukaryotes, and their genomes are much smaller and more simply organized. Despite their simplicity, prokaryotes are hardy survivors. These little guys can be found in such extreme conditions where complex eukaryotic creatures like us and animals cannot even step foot in, such as boiling hot springs, acidic lakes and even radioactive waste! In this article, we will talk about prokaryotic genetics, how genetic variation can arise and genetic regulation.
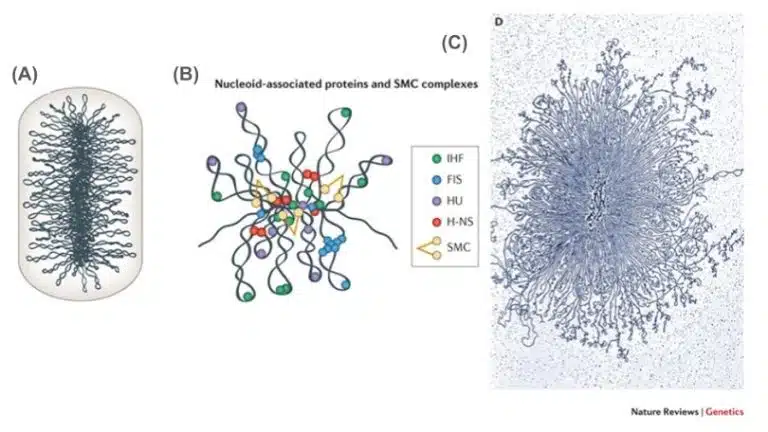
If the eukaryotic genome was described as a Marie Kondo-esque organization for easy storage and retrieval, then the prokaryotic genome can be likened to a single long piece of string that has been crumpled and shoved into a small drawer. Unlike eukaryotes which have multiple linear chromosomes, the main genome of a prokaryotic cell is a single, circular DNA molecule. The prokaryotic genome is also much smaller than a eukaryotic genome (an Escherichia coli cell has 4 to 6 million bases, compared to 3 billion bases in a human), and is located within an area known as the nucleoid region, unlike eukaryotic genomes which are enclosed within the nuclear membrane. The main prokaryotic genome contains all essential genes for survival and reproduction, and is much more compact and efficient compared to an eukaryotic genome, for it does not contain many non-coding regions.
The prokaryotic genome is supercoiled in order to squeeze it within the confines of the cell. Many proteins are involved in maintaining this supercoiled structure. Two important proteins that help to maintain the supercoils are topoisomerase and gyrase. While gyrase introduces supercoils and further compacts DNA, topoisomerase relieves these coils to prevent torsional stress. Together, both proteins maintain the compactness and functionality of the prokaryotic genome.
In addition to the main genome, prokaryotes also carry extra-chromosomal DNA in the form of plasmids. Plasmids are small, circular DNA molecules that exist independently of the main chromosome. Plasmids have their own origin of replication and can replicate itself independently of the main chromosome. The copy number of plasmids may vary; high copy plasmids would replicate prolifically while low copy numbers do not replicate much. Plasmids are commonly involved in horizontal gene transfer, where it is able to be transferred between prokaryotes. These properties make plasmids valuable tools in genetic engineering, where they are widely used as vectors due to their ease of customisation and replication.
While not essential for survival, plasmids often provide extra selective advantages and improve the cell’s survival. Some special abilities that plasmids confer include antibiotic resistance, virulence factors, metabolic enzymes and fertility factors (see conjugation later in this article!)
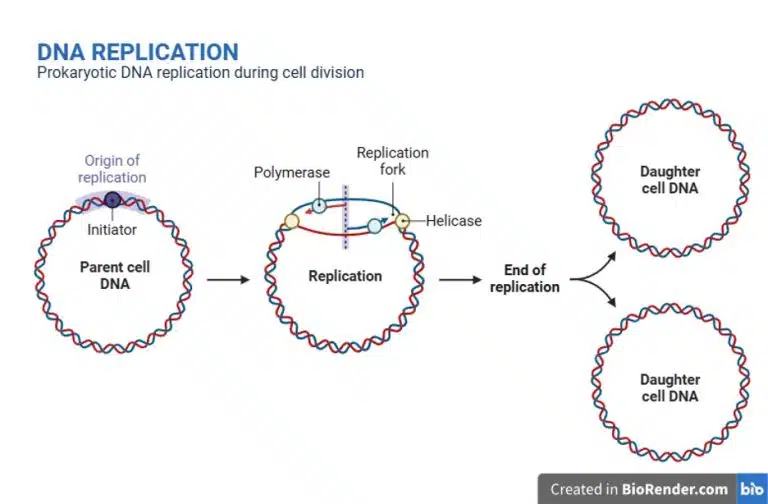
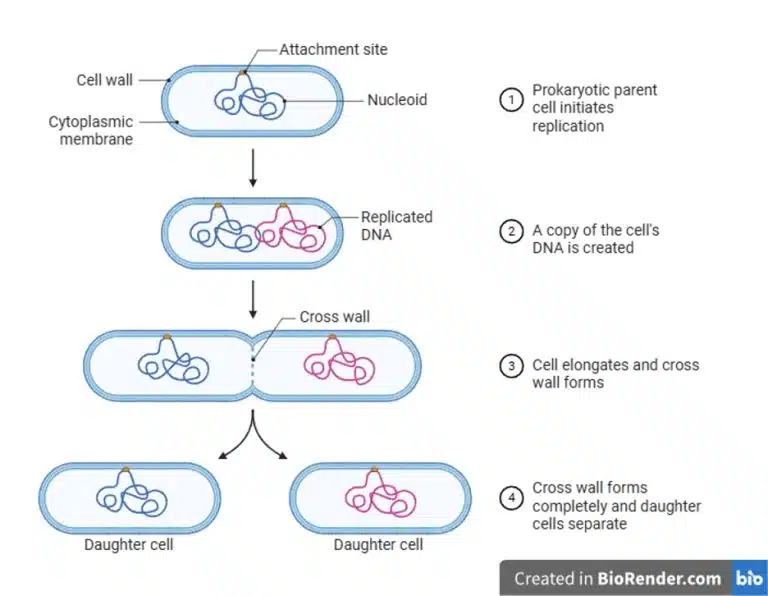
After DNA replication is completed, the prokaryotic cell divides into its daughter cell through binary fission. The cell wall elongates and cross wall forms to demarcate the two daughter cells. After completion of the cross wall, the daughter cells separate as individual prokaryotic cells.
Anyone who has heard of (or even had) an antibiotic resistant bacterial infection would know how sturdy and resilient prokaryotes can be. For us eukaryotes, we get genetic variation through independent assortment and crossing over of homologous chromosomes during meiosis, and also through novel combinations of chromosomes during sexual reproduction. Prokaryotes, on the other hand, reproduce asexually. Yet, they are able to achieve high levels of genetic variation which allows them to adapt rapidly to changing environments.
One method from which genetic variation can arise is through mutations. Errors during DNA replication or exposure to mutagens can result in changes to DNA sequences. If these changes confer beneficial traits to the prokaryote, then these changes are more likely to be maintained and passed down, hence increasing genetic variation.
Bonus information: The E. coli long term evolution experiment is an ongoing study at the University of California. This experiment tracks genetic changes in 12 originally identical populations of asexual E. coli bacteria since 24 February 1998, and has demonstrated how mutations drive genetic variation, which in turn drives natural selection and evolution of the bacteria over the years.
While mutations generate changes in DNA from within the cell, prokaryotes can also acquire new DNA from outside itself through horizontal gene transfer.
During transformation, a prokaryotic cell picks up and incorporates foreign DNA from the environment into its own genome. In nature, cells that are “naturally competent” carry the genes encoding proteins required for the process of DNA transformation, and carry DNA across the cell wall and plasma membranes.
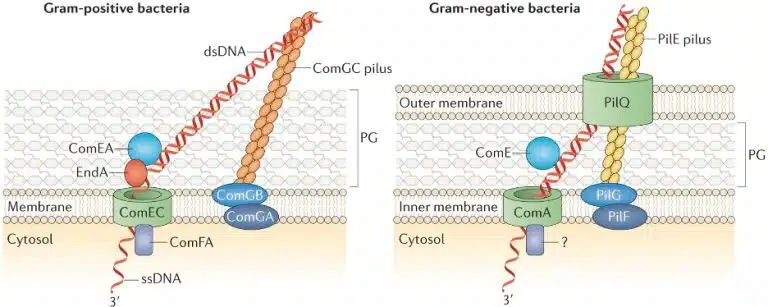
In the laboratory, there are ways to make cells “competent”. This may include treating cells with calcium chloride, or using electric pulses to facilitate the uptake of extracellular DNA (electroporation).
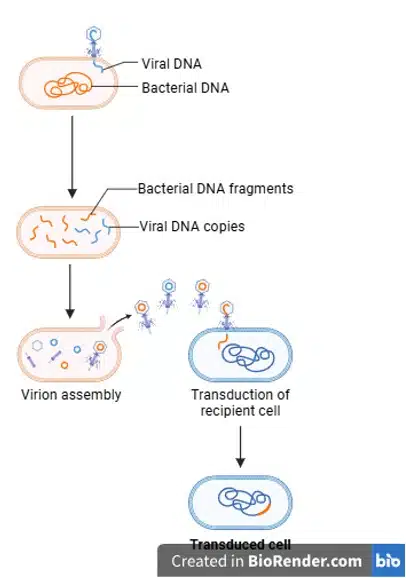
During transduction, DNA from one bacterium is transferred to another through a bacteriophage (bacteria-infecting virus). While in the donor bacteria, the virus may accidentally incorporate some of the donor bacteria’s DNA into itself. When this virus infects the next bacteria, the donor bacteria’s DNA is injected into the recipient bacteria, after which it can be incorporated into the recipient’s genome.
Observe that the drawing only uses simple lines, with no colouring or shading. The only organelles that are visible in the microscopy image are the nucleus and cytoplasm, so those will be clearly labelled. Be sure to replicate the shape and structure of the nucleus accurately, as faithful representation is key to a proper biological drawing.
With that, we have gone through and examined sample questions on the three key components of practical examinations:
Success in the practical exam isn’t just about preparation, it also depends on your focus and attentiveness in the moment. Hence, remember to live in the moment and be observant of experimental outcomes to ace your practical exams! Stay humble, observant and curious!
Find out more by joining us at Science of Studying!
Prepared by: Michelle
During conjugation, DNA is transferred directly from a donor cell to a recipient cell through a physical bridge known as a pilus. The proteins required for pilus generation are encoded within the F plasmid (recall that plasmids can encode fertility factors), and the donor cell carrying the F plasmid is known as an F+ cell. The pilus facilitates the transfer of the F plasmid from the donor to the recipient cell (F– cell). During the transfer, the F plasmid is replicated and the newly synthesized F plasmid is transferred to the recipient, resulting in both cells carrying the F plasmid and becoming F+.
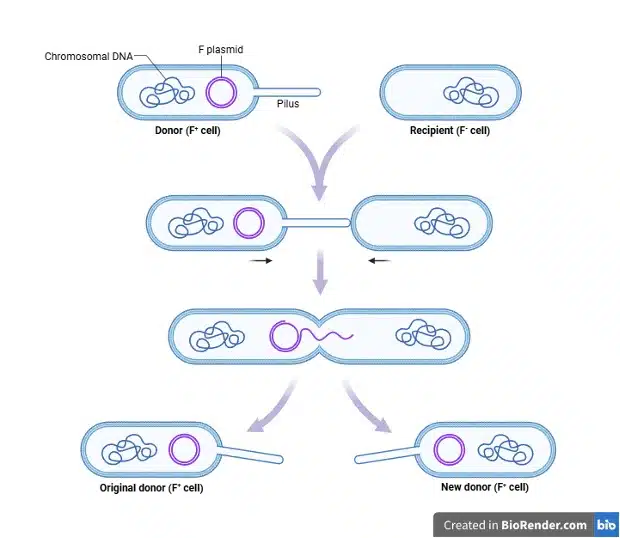
Transformation, transduction and conjugation are all important events that allow a prokaryotic cell to take up extracellular DNA, allowing for exchange of genetic information and adaptation to changing environments. Horizontal gene transfer hence plays a significant role in the spread of traits beneficial to the prokaryote’s survival, such as antibiotic resistance.
After taking up extracellular DNA, the cell would have to incorporate the DNA into its genome. The DNA could end up existing as an independent plasmid (such as the case of the F plasmid), or incorporated into the main genome through homologous recombination or site-specific recombination.
Despite the size and simplicity of the prokaryotic genome, some form of gene regulation still has to take place. Gene regulation allows the prokaryote to use resources even more efficiently, ensuring that genes are switched on (activated) when required or switched off (repressed) when not.
Gene regulation generally takes place at the transcriptional level in prokaryotes, such that the amount of mRNA produced from a gene can be controlled. Remember that transcription and translation usually occur simultaneously in prokaryotes due to the lack of a membrane-bound nucleus. Hence, the amount of mRNA produced directly correlates to the amount of protein produced and the level of gene expression.
Genes that encode proteins with related functions are usually clustered together in operons, which are transcribed together as a single unit. This allows for efficient regulation of multiple genes at once. An operon comes with a regulatory gene which is typically upstream of the operon, which encodes either an enhancer or repressor. The operon itself consists of a promoter, followed by an operator where an enhancer or repressor can bind to either activate or repress transcription. Following the operator would be structural genes that are co-transcribed into a single polycistronic mRNA. These proteins usually function in a common pathway. Finally, the terminator lies at the end of the operon to signal termination of transcription.
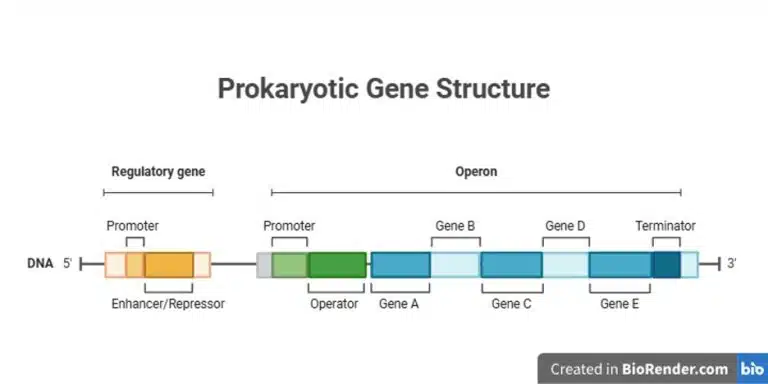
Let us look at two classic examples of operons in E. coli, the lac operon which demonstrates both positive and negative regulation, and the trp operon which is an example of repression.
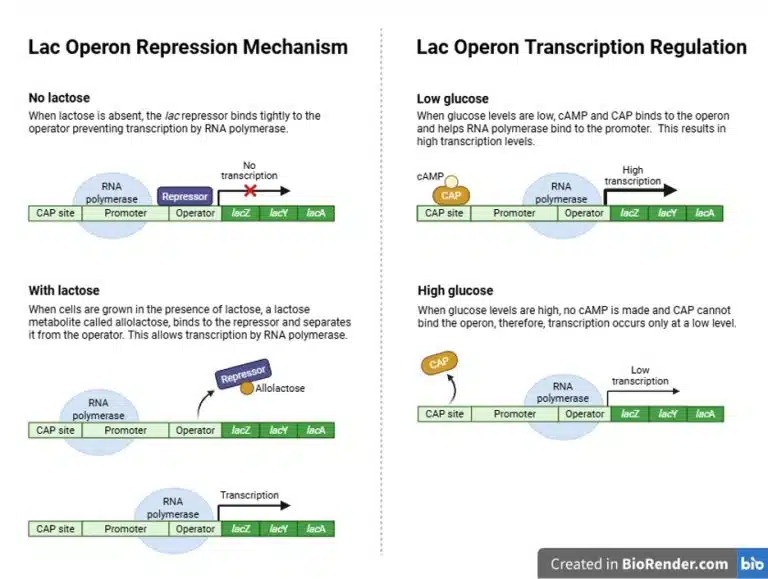
The lac operon enables metabolism of lactose in E. coli. It has a regulatory gene that encodes the repressor protein lacI. Within the lac operon itself, the CAP binding site lies after the promoter, followed by the operator, lacZ (encodes ꞵ-galactosidase), lacY (permease), lacA (transacetylase), and finally the terminator. ꞵ-galactosidase catalyses the breakdown of lactose to galactose and glucose, but also catalyses the conversion of lactose to allolactose. Permease is a transporter protein that allows the uptake of lactose into the cell. The function of transacetylase in the context of lactose metabolism is still not known.
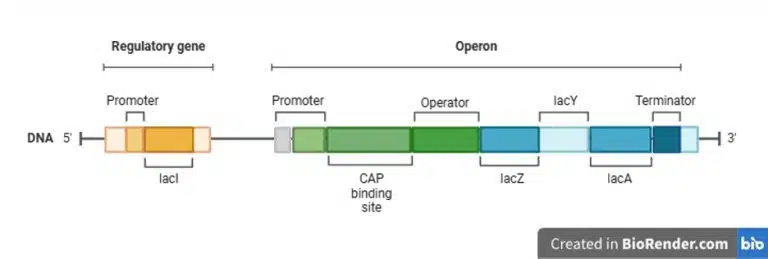
Negative regulation of the lac operon is facilitated by the operator and the repressor protein. The gene lacI encodes the repressor protein, which binds to the operator in the absence of lactose, thus preventing RNA polymerase from transcribing the rest of the structural genes in the operon. However, the binding of repressor to operator is “leaky” and low levels of ꞵ-galactosidase and permease are still produced and present in the cell. In the presence of lactose, permease facilitates the uptake of lactose into the cell, where it is converted into allolactose by ꞵ-galactosidase. Allolactose binds to the repressor protein, inactivating and removing it from the operator.
However, removal of the repressor protein from the operator is not sufficient to activate the lac operon, as RNA polymerase does not bind very well to the promoter on its own. Normally, E. coli would preferably metabolise glucose, even if other sugars are available. The absence of glucose is a signal for positive regulation of the lac operon, which is facilitated by the CAP binding site. When glucose levels are low, cyclic AMP (cAMP) is produced. cAMP then binds to CAP (catabolite activator protein), which then binds to the CAP binding site on the lac operon. The binding of CAP strongly promotes the binding of RNA polymerase to the promoter, thus promoting the expression of the lac operon.
From the example of the lac operon, we can observe both negative and positive regulations being demonstrated. Only in the presence of lactose and absence of glucose, will there be high expression of the lac operon so as to metabolize lactose for the cell’s energy needs.
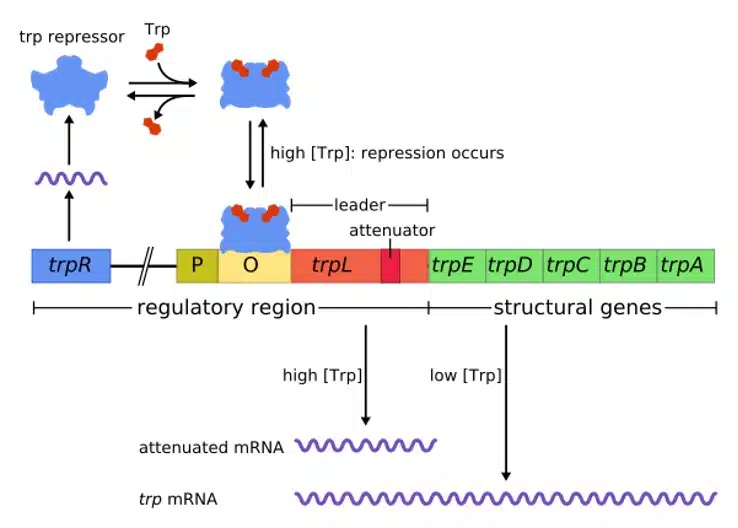
The trp operon is an example of a repressible operon, and allows E. coli to synthesize tryptophan only when needed. The trp operon is regulated by the trpR gene that encodes the trp repressor. The trp repressor is inactive, unless bound by tryptophan. The trp operon consists of the promoter, operator, trpL (leader sequence), the structural genes trpE, trpD, trpC, trpB and trpA, followed by the terminator.
When tryptophan levels are low, the repressor is inactive and unable to bind to the operator, allowing the trp operon to be expressed undisturbed. The expression of structural genes increases tryptophan production, which raises the tryptophan levels within the cell. Once high levels of tryptophan have been reached, there is a higher likelihood that tryptophan binds to its repressor and activates it. The activated repressor binds to the operator, blocking RNA polymerase and repressing the expression of the trp operon. This way, the cell does not waste resources producing more tryptophan when it is already abundant.
Despite their much smaller size and simpler anatomies, prokaryotes are still extremely successful creatures that have survived the test of evolution over millions of years. From streamlined genome organisation to simple and efficient gene regulation, the genetic simplicity of prokaryotes has allowed them to continuously master and adapt to any environment that they are thrown into. This article offers a small glimpse into the broader world of microbiology and genetic engineering. Do get comfortable with the topic of genetics, as it is not only a very testable part of A levels, it is also what generates life from a humble collection of nucleic acids.
Find out more by joining us at Science of Studying!
Prepared by: Michelle
You might want to download a pdf copy of this article for future reference!
Click the white download button below, enter your email, and the pdf file will be delivered to your inbox! (Remember to check spam!)
The Science of Studying provides live online tuition via Zoom classes for Combined/Pure Chemistry, Biology, and Physics. To date, we have taught 800+ students over 12 years.
In case you are wondering, yes – there is a science behind studying!
At Science of Studying, we use our SOS system™ to teach our classes so that even last-minute students can see remarkable improvements in their grades – without mind-numbing memorisation of textbooks and without the drudgery of doing numerous assessment books.
All these conducted in a fun, interactive, stress-free online environment.
If you need help with your Chemistry, Biology, and Physics subjects, do reach out to us and we will see what we can do to help.
Contact Us: Click Here
Admin number: +65 88082348

The SOS system™️ guides students through an effective process of:
Join our proven online tuition programs and see real improvements in understanding, confidence, and school results.
Book a free trial lesson and start the journey today or discover more below:
WhatsApp us